- Home
- About Journals
-
Information for Authors/ReviewersEditorial Policies
Publication Fee
Publication Cycle - Process Flowchart
Online Manuscript Submission and Tracking System
Publishing Ethics and Rectitude
Authorship
Author Benefits
Reviewer Guidelines
Guest Editor Guidelines
Peer Review Workflow
Quick Track Option
Copyediting Services
Bentham Open Membership
Bentham Open Advisory Board
Archiving Policies
Fabricating and Stating False Information
Post Publication Discussions and Corrections
Editorial Management
Advertise With Us
Funding Agencies
Rate List
Kudos
General FAQs
Special Fee Waivers and Discounts
- Contact
- Help
- About Us
- Search

The Open Medical Informatics Journal
(Discontinued)
ISSN: 1874-4311 ― Volume 13, 2019
A Framework for Integration of Heterogeneous Medical Imaging Networks
Carlos Viana-Ferreira*, Luís S Ribeiro , Carlos Costa
Abstract
Medical imaging is increasing its importance in matters of medical diagnosis and in treatment support. Much is due to computers that have revolutionized medical imaging not only in acquisition process but also in the way it is visualized, stored, exchanged and managed. Picture Archiving and Communication Systems (PACS) is an example of how medical imaging takes advantage of computers. To solve problems of interoperability of PACS and medical imaging equipment, the Digital Imaging and Communications in Medicine (DICOM) standard was defined and widely implemented in current solutions. More recently, the need to exchange medical data between distinct institutions resulted in Integrating the Healthcare Enterprise (IHE) initiative that contains a content profile especially conceived for medical imaging exchange: Cross Enterprise Document Sharing for imaging (XDS-i). Moreover, due to application requirements, many solutions developed private networks to support their services. For instance, some applications support enhanced query and retrieve over DICOM objects metadata.
This paper proposes anintegration framework to medical imaging networks that provides protocols interoperability and data federation services. It is an extensible plugin system that supports standard approaches (DICOM and XDS-I), but is also capable of supporting private protocols. The framework is being used in the Dicoogle Open Source PACS.
Article Information
Identifiers and Pagination:
Year: 2014Volume: 8
First Page: 20
Last Page: 32
Publisher Id: TOMINFOJ-8-20
DOI: 10.2174/1874431101408010020
Article History:
Received Date: 19/3/2014Revision Received Date: 9/6/2014
Acceptance Date: 10/6/2014
Electronic publication date: 16 /9/2014
Collection year: 2014
open-access license: This is an open access article licensed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted, non-commercial use, distribution and reproduction in any medium, provided the work is properly cited.
* Address correspondence to this author at the DETI/IEETA, Universidade de Aveiro, Campus Universitário de Santiago, 3810-193 Aveiro, Portugal; E-mail: c.ferreira@ua.pt
| Open Peer Review Details | |||
|---|---|---|---|
| Manuscript submitted on 19-3-2014 |
Original Manuscript | A Framework for Integration of Heterogeneous Medical Imaging Networks | |
INTRODUCTION
Medical imaging systems and networks are valuable tools supporting the medical profession in both decision-making and treatment procedures. Its importance has beengrowing in the last decades, following the tendency of computational resources availability. Technological advances are supporting the emergence of new methods of acquiring medical images, but also new ways of storing, accessing and visualizing data [1Huang H, Ed. PACS and imaging informatics basic principles and applications. Wiley-Blackwell. 2011.]. The major contribute to this reality is the PACS(Picture Archiving and Communication Systems) concept, a term that characterizes systems responsible for the acquisition, storage, visualization and distribution of medical image through a computer network [1Huang H, Ed. PACS and imaging informatics basic principles and applications. Wiley-Blackwell. 2011.-3Costa C, Ferreira C, Bastião L, Ribeiro L, Silva A, Oliveira J. Dicoogle - an Open Source Peer-to-Peer PACS. J Digit Imag 2011; 24(5): 848-56.]. The proliferation of this concept was powered by the DICOM (Digital Imaging and Communications in Medicine) standard [4ACR-NEMA.Digital Imaging and Communications in Medicine (DICOM). In: Rosslyn VA National Electrical Manufacturers Association 2011. ] that ensures the interoperability between equipment’s of different manufactures. DICOM is an extensive and complex standard, defining the network communication layers, the services’ commands, the persistent objects coding, the media exchange structure, the documentation that must follow an implementation, etc. [5Pianykh OS. Digital imaging and communications in medicine (DICOM): a practical introduction and survival guide. Berlin Springer 2008; p: 383.].
PACS are empowering healthcare practitioners with capabilities to access patient’s examinations anytime and anywhere, and having an important role in telemedicine, telework and collaborative work environments. However, those systems also have many opened issues. Firstly, medical imaging repositories are often looked on as “inert bags” ofDICOM objects, accessible only through the DICOM query and retrieve service. Secondly, the increasing adoption of medical imaging equipment in healthcare institutions, including small diagnostic centres, has led to a huge dispersion of data repositories. There is no simple solution for imaging data exchange between multiple places using Web 2.0 Internet, because traditional medical imaging protocols do not perform well in inter-institutional scenarios [6Valente F, Viana-Ferreira C, Costa C, Oliveira JL. A RESTful Image Gateway for Multiple Medical Image Repositories. Information Technology in Biomedcine IEEE Transactions on 2012; 16(3): 356-64., 7Adams SA. Revisiting the online health information reliability debate in the wake of web 2. An inter-disciplinary literature and website review. Int J Med Inform 2010; 79(6): 391-400.]. Typically, bureaucratic and technical arrangements are required, and at the end, the exchange of information is limited to either inside an institution, or among peers in a few well-defined institutions.
However, this scenario is changing. In recent years, a renewed interest in medical imaging networks, with the introduction of state-of-the-art technologies and new paradigms such as Grid, Cloud Computing, Peer-to-Peer networks and indexing engines, has been rising. Moreover, IHE (Integrating Healthcare Enterprise) initiative has defined an integration profile to medical imaging scenario, aiming to promote trans-institutional sharing of data.
Dicoogle is a PACS archive that innovates in the information system and services provided [8Santos M, Bastião L, Costa C, Silva A, Rocha N. DICOM and Clinical Data Mining in a Small Hospital PACS A Pilot Study.ENTER prise Information Systems. In: Cruz-Cunha M, Varajão J, Powell P, Martinho R, Eds. Communications in Computer and Information Science 221. Berlin Heidelberg;: Springer 2011; pp. 254-63.]. It is an open source solution that replaced the traditional relational database by an indexing engine, allowing advanced searches over DICOM objects metadata. The application scenarios are vast like, for instance, research associated to dose surveillance and quality assurance [8Santos M, Bastião L, Costa C, Silva A, Rocha N. DICOM and Clinical Data Mining in a Small Hospital PACS A Pilot Study.ENTER prise Information Systems. In: Cruz-Cunha M, Varajão J, Powell P, Martinho R, Eds. Communications in Computer and Information Science 221. Berlin Heidelberg;: Springer 2011; pp. 254-63.]. To support proprietary services in distributed environments, it was necessary to develop a private network protocol.
This article presents an integrationframework to medical imaging that provides interface with three main protocols: DICOM standard (Storage and Query and Retrieve) to be interoperable with traditional equipment’s of imaging laboratories, a XDS-I to support normalized communications with other institutional domains, and a private communicat-ion protocol to support Dicoogle functionalities in distri-buted environments. Moreover, the solution includes a Data Source Bus that provides data federation services. The bus is capable of receiving a query from a specific network interface and broadcast it to the others interfaces. The query results will be fused in a single list that will be returned to the requester. Finally, the framework also provides an abstraction to transference of DICOM objects between distinct network interfaces. Each member of a network is able to use any of these services, in a transparent way, for searching and also to contribute in a search of other members. The framework proposal will be presented using the Dicoogle project as proof-of-concept.
BACKGROUND
Medical Imaging Laboratories
Medical imaging is a process and technique to acquire visual representations of tissues and organs. Due to the richness of the produced data, this plays, unquestionably, a very important role to support medical diagnosis and treatment [9Sylva P. A Situation Analysis on PACS prospects for a Developing Nation. Sri Lanka J Bio-Med Informa 2010; 1(2): 112-7., 10Rengier F, Mehndiratta A, von Tengg-Kobligk H , et al. 3D printing based on imaging data review of medical applications. Int J CARS 2010; 5(4): 335-41.]. In fact, the importance of medical imaging is still growing. This tendency follows the growth of computational resources availability. Computers are revolutionizing numerous processes related to medical imaging, not only the process of image acquisition itself, but also how the images are stored, visualized and transferred from one place to another.
The concept Picture Archiving and Communication Sys-tems (PACS) embraces a set of systems that are responsible for the storage, acquisition, visualization and distribution of medical imaging data. Initially, these systems were designed to serve only a department of a healthcare institution and, for that reason, they were mainly composed by only the PACS controller and archive, the acquisition equipment and the display workstations, all linked by a local area network which was a very restrictive view of the concept. Soon, PACS were designed to serve the whole healthcare institution and even multiple healthcare institutions. This was the motivation for the telemedicine services development [11Johnson N. Teleradiology 2010 technical and organizational issues. Pediatr Radiol 2010; 40(6): 1052-5., 12Klymash MM, Romanchuk VL, Krasko OV, Eds. Telemedicine system model using Internet-technology. Modern Problems of Radio Engineering Telecommunications and Computer Science (TCSET) 2010 International Conference on 2010 23-27 Feb. 2010.]. Nevertheless, there was a critical issue in combining the systems of different departments: systems of different manufacturers were not interoperable, because each manufacturer had a file format to store the files, a protocol of communications, and so on.
Digital Imaging and Communications in Medicine
Digital Imaging and Communications in Medicine (DICOM) [4ACR-NEMA.Digital Imaging and Communications in Medicine (DICOM). In: Rosslyn VA National Electrical Manufacturers Association 2011. ] (currently in version 3) resulted from an effort to standardize the processes of a PACS. Among others, this standard defines: the network communication layers, the services commands, the persistent objects coding, the media exchange structure and the documentation that must follow an implementation [4ACR-NEMA.Digital Imaging and Communications in Medicine (DICOM). In: Rosslyn VA National Electrical Manufacturers Association 2011. ].
As for communications, DICOM defines which and how information will be exchanged in two steps. Firstly, there must be a negotiation phase, where numerous relevant aspects are agreed upon. For instance, the image modality that will be transmitted (x-ray angiography, Ultrasound, computed tomography, etc.) and how this data will be encoded (little endian or big endian, uncompressed or compressed, and so on). Secondly, if the negotiation process is successfully completed, then effective data transfer between the two hosts takes place.
The part 5 of the DICOM standard defines the structure and encoding of a DICOM document. Basically, a DICOM document is formed by a set of Data Elements, for instance: pixel image data, free-text, structured reports, image modality, information about the patient, equipment’s reference, acquisition parameters, image resolution [4ACR-NEMA.Digital Imaging and Communications in Medicine (DICOM). In: Rosslyn VA National Electrical Manufacturers Association 2011. ]. The standard defines some of them as mandatory, while others are only optional, accordingly to the modality specifications [13Lemoigne Y, Caner A Eds, Gibaud B, Eds. The DICOM Standard A Brief Overview. In Molecular Imaging Computer Reconstruction and Practice NATO Science for Peace and Security Series B Physics and Biophysics Springer Netherlands. 2008; pp. 229-38.].
Data dealt by healthcare institution are privacy demanding. For that reason, these institutions define in their firewalls, very restrictive policies that allow only traffic of “well behaved” protocols [6Valente F, Viana-Ferreira C, Costa C, Oliveira JL. A RESTful Image Gateway for Multiple Medical Image Repositories. Information Technology in Biomedcine IEEE Transactions on 2012; 16(3): 356-64.] like HTTP (Hypertext Transfer Protocol) and IMAP (Internet Message Access Protocol). Even DICOM communication protocol messages are usually blocked by these restrictive policies. This problem was minimized by the 18th part of the standard that defines the Web Access to DICOM objects (WADO). It is oriented to the distribution of images or other medical data via web [4ACR-NEMA.Digital Imaging and Communications in Medicine (DICOM). In: Rosslyn VA National Electrical Manufacturers Association 2011. ]. This protocol allows the remote access to DICOM files through HTTP or HTTPS (HTTP Secure) channels. Basically, with an implementation of this part of the standard, the client sends an HTTP Request (GET) with a URL/URI of the wanted object. DICOM defines the response as a HTTP Response whose message body is encoded in one of the following formats: DICOM, JPEG, GIF, PNG, JP2, MPEG, plain text and HTML. Besides, it is also recommended that the server supports XML, PDF and RTF as well, not discarding the possibility to support other file formats.
WADO is very simple to implement and use, giving the possibility of receiving DICOM objects, without the need to “speak” DICOM. The client has to specify the DICOM wanted object through its unique identifiers. In fact, WADO solves one of the issues of the communications from the outside to the inside of an institution, which are the firewalls restrictive policies of healthcare institutions. Therefore, as WADO uses HTTP or HTTPS as the communication protocol, usually the firewalls allows that traffic. Nevertheless, this part of the DICOM standard does not support any search mechanism. The files are only retrieved if the client sends the unique identifier of the document, which means that the client must know it in advance.
In resume, DICOM is a well-accepted standard being implemented in most systems of this kind. Actually, nowadays, almost all medical imaging equipment manufacturers provide embedded DICOM interfaces in their products. This solved most of the interoperability problems inside institutions. However, some problems concerning interoperability between systems of distinct institutions are not solved by this standard.
Integrating Healthcare Enterprise
Numerous problems related with the interoperability between systems of healthcare institutions were solved by DICOM standard. Nevertheless, some problems still rem-ained concerning interoperability between systems of distinct healthcare institutions. As such, collaboration between healthcare enterprise professionals resulted in the Integrating Healthcare Enterprise (IHE) initiative, which aimed to the promotion of collaboration between institutions [14Teng C C, Mitchell J, Walker C, Swan A, Davila C, Howard D , Eds. A medical image archive solution in the cloud. Software Engineering and Service Sciences (ICSESS) 2010 IEEE International Conference on 2010 IEEE. ]. In fact, IHE is not a standard per se, instead, it is a meta-standard that defines how existing standards should be used, defining integration profiles. These integration profiles provide the guidelines for building integration ready systems. Each profile is composed by several actors and transactions. The IHE actors are abstractions of real-world systems and transactions the inter-actor communication abstractions.
Cross-Enterprise Document Sharing for Imaging
Cross-Enterprise Document Sharing [15IHE.IHE IT Infrastructure (ITI) Technical Framework Revision 10.0 Final Text. IHE 2013. In: ] (XDS) integration profile provides core guidelines for sharing documents among all healthcare institutions, more precisely, supporting querying, retrieving, publishing and registering EHR documents.
XDS for Imaging (XDS-I) [16Seifert P. IHE Radiology Technical Framework Supplement XDS-I. Integration Profile 2009 ] is a content profile scoped to the medical imaging domain, where systems like PACS, RIS or DICOM objects are taken into account on the XDS architecture. Fig. (1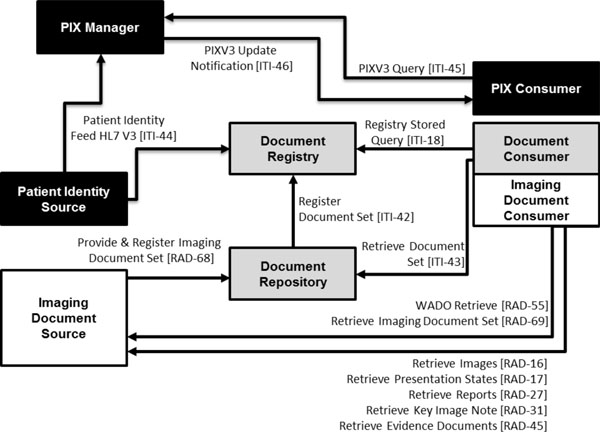 ) shows the actors taking part in the XDS-I profile and the transactions between actors. The Imaging Document Source is the actor that provides, or holds, the clinical documents (e.g. PACS). It is responsible for pushing documents and their metadata to the respective Document Repository. For DICOM objects, the Repository will only hold the DICOM key object selection (KOS). KOS objects are small DICOM objects containing a list of UID references (instead of the image data itself) that enables the Document Consumer to retrieve the images from the Imaging Document Sources. Besides being responsible for persistent documents, the Repository actor also registers the document (forwarding its metadata) in the appropriate Registry. The Registry actor is the central player of the entire XDS affinity domain. It maintains the metadata, which enables cross-organization document discovery. This metadata, that describes the document entitle XDS Document Entry, is a specialization of the Extrinsic Object of the ebXML RIM standard developed by OASIS1. The Extrinsic Object is an ensemble of other ebRIM objects, such as Slot or Classification objects. The Registry responds to the consumer’s queries retrieving the XDS document entries that fit the query. With this information, the consumer contacts the respective Imaging Document Source requesting access to the DICOM object(s). If access is granted, the set of images is then typically retrieved through the WADO protocol.
) shows the actors taking part in the XDS-I profile and the transactions between actors. The Imaging Document Source is the actor that provides, or holds, the clinical documents (e.g. PACS). It is responsible for pushing documents and their metadata to the respective Document Repository. For DICOM objects, the Repository will only hold the DICOM key object selection (KOS). KOS objects are small DICOM objects containing a list of UID references (instead of the image data itself) that enables the Document Consumer to retrieve the images from the Imaging Document Sources. Besides being responsible for persistent documents, the Repository actor also registers the document (forwarding its metadata) in the appropriate Registry. The Registry actor is the central player of the entire XDS affinity domain. It maintains the metadata, which enables cross-organization document discovery. This metadata, that describes the document entitle XDS Document Entry, is a specialization of the Extrinsic Object of the ebXML RIM standard developed by OASIS1. The Extrinsic Object is an ensemble of other ebRIM objects, such as Slot or Classification objects. The Registry responds to the consumer’s queries retrieving the XDS document entries that fit the query. With this information, the consumer contacts the respective Imaging Document Source requesting access to the DICOM object(s). If access is granted, the set of images is then typically retrieved through the WADO protocol.
 |
Fig. (1) Combined diagram of the PIX and XDS-I profiles. |
 |
Fig. (2) Framework plugin architecture. |
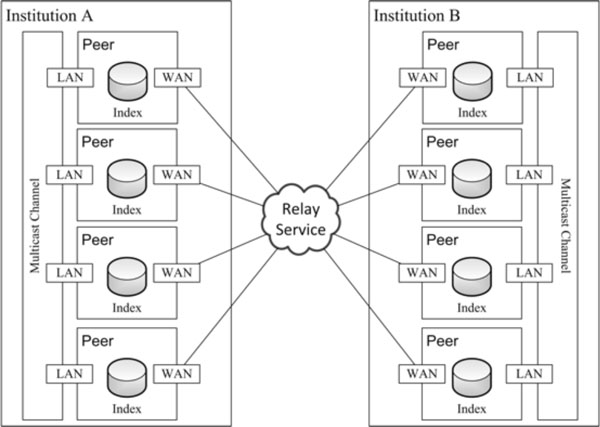 |
Fig. (4) P2P network architecture. |
 |
Fig. (5) Relay installer software. |
 |
Fig. (6) Messages examples: query request, query response and file request. |
 |
Fig. (7) Dicoogle desktop interface. |
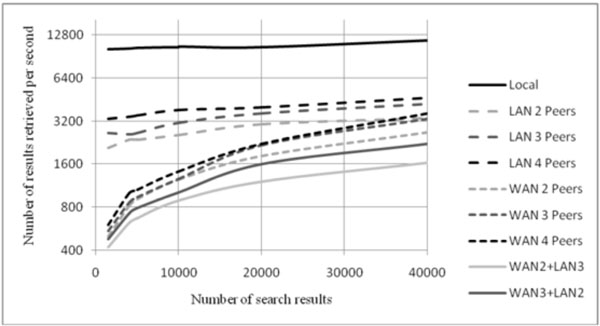 |
Fig. (8) Graph of the number of results retrieved per second as a function of the total number of search hits. |
 |
Fig. (9) Test scenarios in which more than one network protocol is used. |
Patient Identifier Cross-Referencing
Patient Identifier Cross-Referencing (PIX) integration profile [17IHE.Patient Identifier Cross-Reference HL7 V3 (PIX V3):and Patient Demographic Query HL7 V3 (PDQ V3). IHE Internatonal 2009.] supports cross-referencing of the patient’s identifiers from multiple institutions. Typically, each patient has a unique identifier within each healthcare institution (the institution’s patient identifier domain), relating its demographic data with its medical records. Therefore, a patient in an inter-institutional scenario will have, at least, an identifier per institution. The multiplicity of identifiers issue is addressed by the PIX profile. It correlates the information of a single patient from the multiple sites where the patient has medical records, enabling a complete view of the patient’s documents within the affinity domain. Fig. (1 ) shows the actors and the transactions (HL7 v3 messages) involved in this profile. The Patient Identity Source (e.g. RIS, HIS) is responsible for feeding patient identifiers of independent domains to the PIX Manager. In its turn, the PIX Manager is typically a central and unique entity that is in charge of creating, maintaining and providing a patient-centric list of identifiers, correlating each entry with the respective patient domain source. This way it is possible to map all the patient identifiers among the different domains. Finally, the PIX Consumer may query the manager actor and, as consequence, discover the patient identifier of the respective domain.
) shows the actors and the transactions (HL7 v3 messages) involved in this profile. The Patient Identity Source (e.g. RIS, HIS) is responsible for feeding patient identifiers of independent domains to the PIX Manager. In its turn, the PIX Manager is typically a central and unique entity that is in charge of creating, maintaining and providing a patient-centric list of identifiers, correlating each entry with the respective patient domain source. This way it is possible to map all the patient identifiers among the different domains. Finally, the PIX Consumer may query the manager actor and, as consequence, discover the patient identifier of the respective domain.
Dicoogle
Dicoogle PACS (http://www.dicoogle.com) is a free and open source solution that supports medical imaging storage and DICOM data mining. It allows the extraction of information from the PACS image archives and performs flexible queries on DICOM metadata [3Costa C, Ferreira C, Bastião L, Ribeiro L, Silva A, Oliveira J. Dicoogle - an Open Source Peer-to-Peer PACS. J Digit Imag 2011; 24(5): 848-56.]. The software installation requirements are minimal and their flexibility to extract different level of data is high. It can run as operating system service but also as a portable application. So, the utilization of this software, as a knowledge extraction tool, only needs to have a view over repository file-system, with read-only permission.
The data produced by digital modalities generate information on performed exams that can be used in different scenarios, like, for example, to monitor patient radiation doses, radiographic procedures and image quality. Some important parameters such as image processing parameters, exposure index, patient dose and geometric information are generated by the modality and transferred to the PACS repository. Despite of this information being stored on the repository imaging objects, the information systems are not able to access them.
The flexibility to index different DICOM objects with different data elements is due to the replacement of the traditional database by an indexing engine. With this approach, it is possible to index all existent DICOM data elements (text-based) without the need to create new fields, new tables, and new relations that would be necessary in the database supported approach. Dicoogle index engine is based on Apache Lucene [18Apache S. Lucene Index Server 2011 cited 2011. Available from http //luceneapacheorg ], a Java index and search library that is used in a large variety of applications, including Wikipedia. It explores the two different types of indexing features supported by Lucene: full text and metadata; and allows the realization of two types of data indexing: a hierarchical content indexing of the DICOM metadata (patient, study, series, image) and a text content indexing (free text query) [19Costa C, Freitas F, Pereira M, Silva A, Oliveira JL. Indexing and retrieving DICOM data in disperse and unstructured archives. Int J CARS 2009; 4(1): 71-7.]. Optionally, it is also possible to extract an image thumbnail, with a configurable matrix, that is saved with indexes. The initial indexing process is a very intensive task than can take days or weeks to finish, depending on repository number of DICOM objects. However, Dicoogle allows us to adjust this process intensity to not perturb regular PACS operation.
The main reason to use Dicoogle is its capacity to query any DICOM attribute, in a “google like” way, and one single query search can include one or more attribute fields (Fig. 1 ). For instance, to search RF modality studies with the text “PULSED” in the “Radiation Mode” attribute field, the following query should be used: [Modality: RF AND Radiation Mode: PULSED]. It also enables to analyse DICOM attributes values pre-established by the user and export the data in Excel format, making possible the indexed data post-processing, (e.g. perform quantitative analysis over the retrieved data).
). For instance, to search RF modality studies with the text “PULSED” in the “Radiation Mode” attribute field, the following query should be used: [Modality: RF AND Radiation Mode: PULSED]. It also enables to analyse DICOM attributes values pre-established by the user and export the data in Excel format, making possible the indexed data post-processing, (e.g. perform quantitative analysis over the retrieved data).
Dicoogle was already used in the real healthcare environment. For instance, in [8Santos M, Bastião L, Costa C, Silva A, Rocha N. DICOM and Clinical Data Mining in a Small Hospital PACS A Pilot Study.ENTER prise Information Systems. In: Cruz-Cunha M, Varajão J, Powell P, Martinho R, Eds. Communications in Computer and Information Science 221. Berlin Heidelberg;: Springer 2011; pp. 254-63.], Dicoogle identified some data inconsistencies and an increase of X-ray exposure level in mammographic studies, performed in a 3 years interval time, using specific queries and data statistical post-processing.
Related Work: Inter-Institutional Communications
Nowadays, the inter-institutional exchange of imaging data is done through traditional channels, including mail, email, patient or private solutions over virtual private networks (VPN). However, all these solutions have drawbacks that can make data exchange infeasible. For instance: (1) VPN can be hard to set up and manage for bureaucratic and technical reasons; (2) traditional mail can take too long; (3) the patient can forget, lose or damage the exams; (4) through email it is necessary to know, previously, where the data will be needed, or to wait until the person responsible for the data answers the request. Thus, efforts have been made to solve this problem. Therefore, the need to develop a system with a distributed repository has been addressed in the literature, in recent years.
DIPACS [20Onbay TU, Kantarci A. Design and implementation of a.distributed teleradiaography system DIPACS 2011; 104(2): 235-42.] is a Peer-to-Peer (P2P) tele-radiography system. This system uses Java Remote Method Invocation (RMI) for communication between different institutions. Nevertheless, the architecture of this system raises some drawbacks, such as the fact that each institution needs to have a gateway that communicates via RMI to the gateways of other institutions. Also, this system needs the setup of routers and firewalls to allow the communication via RMI. Moreover, it does not have an automatic discovery mechanism: the nodes of the system must be previously registered in a server in the Internet.
In [21Bian J, Seker R, Topaloglu U, Eds. A Secure Distributed File System for Medical Image Archiving. Social Computing (SocialCom) 2010 IEEE Second International Conference on 2010 IEEE. ], Bian et al. described a distributed file system based on P2P paradigm. This system is especially focused on the distribution and security of data among the nodes that belong to the system. However, the node identification and discovery is carried out through broadcast of messages, which is not possible in Internet. Therefore, the nodes of this system are only able to communicate with each other, if they are inside the same local area network.
Other P2P systems were also developed, such as the ones described in [22Blanquer I, Hernández V, Mas F, Eds. A p2p platform for sharing radiological images and diagnoses. Proc DiDaMIC Workshop 2004 Citeseer. , 23Teles CP, Castro H , Eds. JMED Uma arquitetura de sistemas Peer-to-Peer para services de assistência médica distribuida à distância. X Congresso Brasileiro de Informática em Saúde Florianópolis Anais do X CBIS 2006. ]. Those two systems are based on JXTA, and both use the rendezvous peer for promoting communications between peers of different institutions. Nevertheless, none of those systems provide an easy to use mechanism to facilitate the setup of the system. Besides, the use of only one node as rendezvous compromises the scalability of the system.
Solutions based on grid paradigm are also present in literature, as the ones described in [24Breton V, Medina R, Montagnat J. DataGrid, prototype of a biomedical grid. Met Inform Med 2003; 42(2): 143-7.-29Vossberg M, Tolxdorff T, Krefting D, Eds. DICOM image communication in globus-based medical grids. Information Technology in Biomed cine IEEE Trans. 2008; 12: pp. (2) 145-53.]. One of the greatest disadvantages of those systems are the bureaucratic and technical issues related to join a grid [30Koutelakis GV, Lymperopoulos DK , Eds. A Grid PACS Architecture Providing Data-centric Applications through a Grid Infrastructure. Engineering in Medicine and Biology Society 2007 EMBS 2007 29th Annual International Conference of the IEEE; 2007 IEEE. ].
With the advent of the cloud, some systems were also developed to take advantage of such technology, like the ones described in [14Teng C C, Mitchell J, Walker C, Swan A, Davila C, Howard D , Eds. A medical image archive solution in the cloud. Software Engineering and Service Sciences (ICSESS) 2010 IEEE International Conference on 2010 IEEE. , 31Silva LAB, Costa C, Oliveira JL. A PACS archive architecture supported on cloud services. Int J CARS 2012; 7(3): 349-58.]. In these two examples, the cloud is used to maintain a central repository of the medical imaging data. In this way, the repository is endowed with the scalability and resiliency inherent in the cloud technology. However, moving the repository from the institution to the cloud raises latency problems, since the velocity to access data stored somewhere in Internet is usually slower than accessing data stored inside the same institution. Another example of the use of cloud is the system described in [32Hsieh J-c, Hsu M-W. A cloud computing based 12-lead ECG telemedicine service. BMC Med Inform Decis Mak 2012; 12(1): 1-12.], where Hsieh and Hsu used cloud to support an ubiquitous system for 12-lead ECG with the possibility of accessing the data through a mobile device.
In conclusion, the literature lacks of a system for medical imaging sharing with the following characteristics: (1) easy to deploy and maintain; (2) able to grow accordingly to the needs; (3) that takes advantage of the reduced latency of Local Area Network (LAN) environment to retrieve data that is already stored inside the institution. The problem of latency is one of the most critical in this kind of systems, as described in [33Langer SG, French T, Segovis C. TCP/IP optimization over wide area networks implications for teleradiology. J Digit Imag 2011; 24(2): 314-21.], since Wide Area Network (WAN) is a high latency environment.
INTEGRATION FRAMEWORK - PROPOSAL
As expressed, Dicoogle will be used as proof-of-concept of proposed integration framework. So, the proposed will be presented using this challenging context.
The main architectural advantage of proposed integration framework is its extensibility supported by a plugin mechanism that promotes the development of new features, in an easy and simple way (Fig. 2 ). This flexibility will be used to support distinct medical imaging network interfaces (i.e. services).
). This flexibility will be used to support distinct medical imaging network interfaces (i.e. services).
The network interfacing is fundamental to permit interoperability with third systems. Moreover, Dicoogle was special requirements, namely the support of enhanced queries over distributed medical imaging environments. So, our architecture must contemplate two distinct layers of communications, described in detail in the next sub-sections:
- Standard: It is an indispensable communications module that ensures standard interactions with other equipment’s through DICOM and XDS-I.
- Dicoogle Proprietary: It is a private network developed to support enhanced search and retrieve of imaging studies inside a farm of Dicoogle peers, i.e. supports a federated view of repositories. Without this layer, Dicoogle searching and data-mining are not available in distributed environments. This network specification follows the peer-to-peer paradigm and uses a relay service hosted at Cloud.
The Data Source Bus (Fig. 2 ) is the component of the architecture responsible for the communication between the communication interfaces of each instance. Thanks to this, when an application instance receives a request from any network interface, it can forward it to the other interfaces in order to answer the request with results from multiple sources: LAN, WAN, XDS-I and DICOM. Moreover, the framework supports also interface with other local data sources. For instance, it was developed a plugin to Dicoogle database, i.e. the Lucene Index Engine (Fig. 2
) is the component of the architecture responsible for the communication between the communication interfaces of each instance. Thanks to this, when an application instance receives a request from any network interface, it can forward it to the other interfaces in order to answer the request with results from multiple sources: LAN, WAN, XDS-I and DICOM. Moreover, the framework supports also interface with other local data sources. For instance, it was developed a plugin to Dicoogle database, i.e. the Lucene Index Engine (Fig. 2 ). In this way, the system is able to give as much data possible in a transparent way.
). In this way, the system is able to give as much data possible in a transparent way.
Standard Interface
DICOM
The framework DICOM networking functionalities are supported by DCM4CHE [27Luo Y, Jiang L, Zhuang T-g. A grid-based model for integration of distributed medical databases. J Digit Imag 2009; 22(6): 579-88., 28Ni Y-J, Youn C-H, Song H, Kim B-J, Han Y, Eds. A PACS-Grid for Advanced Medical Services based on PQRM. Intelligent Sensors Sensor Networks and Information 2007 ISSNIP 2007 3rd International Conference on 2007 IEEE. ]. This SDK is used to extract DICOM data elements from persistent objects and to implement the Storage SCP (Service Class Provider) and Query/Retrieve SCP services. In the Storage process, the archive receives a C-STORE request for each DICOM object (e.g. images). In the Dicoogle case, the objects are stored on the repository file-system and are also analysed by a DICOM metadata extractor. All information extracted is stored in an index engine system, namely the Apache Lucene. The Dicoogle administrator can configure several aspects like, for instance, define the level of information extracted from DICOM objects metadata: only DIM (DICOM Information Model) mandatory fields, all DICOM object attributes and all (or part of) mandatory fields from specific IOD (Infor-mation Object Definition) image modality. So, it is possible to have distinct configurations according to medical imaging modality.
In a DICOM query process a specific Boolean expres-sion, using the indexed DIM fields, is applied. Each C-FIND Request (i.e., the DICOM query command) [34ACR-NEMA.Digital Imaging and Communications in Medicine. Part 7 Message Exchange 2011 ], received by Dicoogle DICOM interface, is mapped to a Dicoogle query using specific keywords such as Patient Name, Study Date, Modality, or other DIM fields. For instance, “Patient Name: FELIX* AND Study Date: [20090101 TO 20090131] AND Modality: CT”. This query is send to the Lucene index engine. The query results will include the location of DICOM persistent objects and all indexed DICOM tags. The fields are selected to build an answer compliant with DIM model. For instance, if C-FIND Request level is Study, it creates a list of studies filtered by Study Instance UID with a structure containing the study and patient mandatory fields for C-FIND Response.
The DICOM retrieve service is based on C-MOVE command [34ACR-NEMA.Digital Imaging and Communications in Medicine. Part 7 Message Exchange 2011 ]. Dicoogle uses the Study Instance UID keyword to interrogate the indexer. If the study exists, it will be returned to the locations of DICOM objects. Those files will be sent to the remote DICOM host, i.e. the client.
XDS-I
In order to capacitate framework clients with standard inter-institutional functionalities, we developed the XDS-I plugin. XDS-I interface implements the following XDS/XDS-I transactions: Provide and Register Imaging Document Set (RAD-68), Retrieve Document Set (ITI-43), Registry Stored Query (ITI-18), and WADO retrieve (RAD-55 –previously provided by Dicoogle core platform). We used SOAP 1.2 as XDS (part B) imposes and for generating the respective stubs for the web-service consumers and the skeletons for the web-service providers we used IHE’s WSDL2 with the API JAX-WS 2.6. The security model follows an SSL/TLS with client X.509 certificates for authentication, in accordance with the Audit Trail Node Authentication (ATNA) directives (transaction ITI-19) providing authentication, channel confidentiality and integrity. After installation, certificates are used on the SSL/TLS handshake to exchange the session symmetric key to secure the communication channel between the IHE actors.
Fig. (3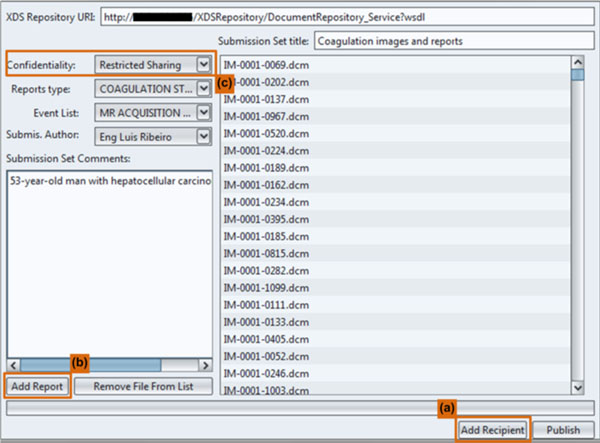 ) shows a screenshot of the graphic user interface for this plugin. In this screenshot, among others, user can choose which images should be shared and with whom, setting the confidentiality level.
) shows a screenshot of the graphic user interface for this plugin. In this screenshot, among others, user can choose which images should be shared and with whom, setting the confidentiality level.
Dicoogle Peer-To-Peer Network
Dicoogle solution allows, not only the access to DICOM persistent objects locally stored, but also provides a transparent view of remote repositories that are linked by a Peer-to-Peer network. In this section, it is described the protocol and topology of this network that allows the search and retrieval of DICOM files.
The proposed system differs from the traditional P2P file sharing systems, since this is not designed to support a huge number of peers. Nevertheless, the number of files in each peer is expected to achieve the order of magnitude of thousands, or even millions, of files. For this reason, the strategy chosen was making each peer responsible for the query and retrieve of their own files. Therefore, each peer is endowed with a local index that maintains a registry of its DICOM documents [19Costa C, Freitas F, Pereira M, Silva A, Oliveira JL. Indexing and retrieving DICOM data in disperse and unstructured archives. Int J CARS 2009; 4(1): 71-7.] (Fig. 4 ).
).
Architecture
P2P is a paradigm of distributed systems where each participant (i.e. peer) shares a part of their own resources. These resources may be files, CPU cycles, storage, printers, etc. In this way, a peer-to-peer system is able to provide a service composed by the set of resources that all peers share [35Schollmeier R. A definition of peer-to-peer networking for the classification of peer-to-peer architectures and applications. Peer-to-Peer Computing 2001 Aug 2001 ]. This paradigm is commonly associated with file sharing applications, since it is one of the areas where this technology is most widely used. However, it is used in numerous scenarios such as instant messaging, Voice over IP, software publication and distribution, search engines, backup services, multimedia streaming, storage services and multiplayer online gaming.
Such distributed systems usually have two main challenges: (1) the bootstrap problem, in other words, how a peer discovers the network and (2) how to search for resources. To overcome these challenges, the P2P paradigm has been changing. In the beginning, a server maintained a list of shared resources. In this way, this server was not only the solution for the searching challenge, but also the solution for the bootstrap problem, since it was the fixed point in Internet where peers may communicate for joining the network. Afterwards, the server was discarded from the P2P system architecture and the queries were flooded through the network instead. Currently, to solve this issue the solution is the usage of Distributed Hash Tables (DHT). However, DHT is not suitable for our scenario, i.e. a distributed set of medical imaging repositories with millions of DICOM documents, since the cost of rearranging the structure when a peer joins or leaves the network would be unaffordable.
After analysing medical imaging workflows, especially the inter-institutional processes, we decided to avoid the creation of unified indexes of resource, being each peer responsible for maintaining only an index of its own DICOM repository. This strategy reduces the network overloading when one peer (with numerous DICOM documents) joins the network. On other hand, it obligates to contact all peers to process a search.
The proposed P2P architecture (Fig. 4 ) is supported by three data acquisition modules, developed as plugins, which interact between them through Framework Datasource Bus API (Fig. 2
) is supported by three data acquisition modules, developed as plugins, which interact between them through Framework Datasource Bus API (Fig. 2 ):
):
- Index Engine – to support local indexing of peer resources, i.e. DICOM meta-data;
- Reliable Multicast – to support Dicoogle communications inside Intranet;
- Relay Service – to support Dicoogle communication in Internet.
Reliable Multicast Communications
Concerning the Dicoogle peers communication in Intranet, it was chosen a fully decentralized network topology, minimizing the infrastructure requirements and the points of failure. Another advantage of this topology is the stability of the network when composed by a small number of peers. The proposed LAN module is based on Jgroups [36Ban B. JGroups a toolkit for reliable multicast communication 2002. Available from http //wwwjgroupsorg ] and uses reliable multicast communications. The Jgroups concept of group was extended to support Dicoogle affinity domains and a new network message protocol was created to support medical imaging services.
To join a group, i.e. an affinity domain, a peer sends a multicast message with the name of the group it wants to join. If the group already exists, one group member will return a list containing the active peers. If not, a new group is created with that single peer. After associated to a group, a peer can send Dicoogle proprietary messages to all peers or to a particular node, using Jgroup communications channels. Using this communication platform, we can assure that all Dicoogle messages are effectively sent and received by destination peers. Finally, when a peer leaves the group, all remaining peers are notified.
Relay Service
Healthcare institution networks are usually not prepared to support communications from the outside to the inside of institutions and, even from the inside to the outside, communications are limited to a reduced set of protocols and ports. So, the implementation of new services obligates, many times, to adjust network configurations or to establish dedicated communication channels (e.g. VPN), consuming time and resources.
To avoid the previously mentioned constraints, we decided to implement an inter-institutional communications module fully compliant with Web 2.0. Taking into consideration that a peer located inside a network institution can only send requests and receive responses, we decided to use a Relay service to bypass this limitation. This bridge unit forwards messages to their destinations, transforming requests into responses. Moreover, the system uses HTTPS protocol to support communications between Dicoogle peers and Relay server.
This Relay is a key element in communications and must be stable, resilient and scalable. The option was to design a software module of easy instantiation at Cloud, namely by institutions using Dicoogle. After analysing the services offered by Cloud providers, we decided to use a Cloud PaaS (Platform as a Service) solution and develop the service relay as a web application. The Dicoogle Relay service was developed to run in Google App Engine (GAE) [37Viana-Ferreira C, Costa C, Oliveira JL, Eds. Dicoogle relay - a cloud communications bridge for medical imaging. Computer-Based Medical Systems (CBMS) 2012 25th International Symposium on 2012 20-22 June 2012. ]. This choice does not require the acquisition and setup of a virtual machine (i.e. an operating system) and has several advantages like, for instance, the automatic scaling, the service resiliency and the geo-distribution of application platform. The automatic scaling is a very important feature to our scenario, because the state of Dicoogle peers can change very quickly from idle to very active, generating significant amounts of traffic.
Finally, a software utility (i.e. Relay Installer – Fig. 5 ) was developed to facilitate the setup of new services. The administrator of Dicoogle only needs to create an account in the GAE platform and use the Installer to setup the relay. The new service has an address that must be configured in every Dicoogle peer associated to this new domain.
) was developed to facilitate the setup of new services. The administrator of Dicoogle only needs to create an account in the GAE platform and use the Installer to setup the relay. The new service has an address that must be configured in every Dicoogle peer associated to this new domain.
Messages and Workflows
As expressed, it was necessary to specify and instantiate a new network message protocol to support medical imaging services. Those messages are exchanged between Dicoogle peers and are classified into four types: Query Request, Query Response, File Request and File Response. The first three are XML-based messages (Fig. 6 ) and the last one is binary.
) and the last one is binary.
The search process begins when a peer (X) dispatches a Query Request message, through multicast or relay service, to one specific target or to all peers of the group. The destination peer receives this message and performs the search on their other plugins. If query resultsexist, a Query Response Message will be used to return them to peer X. This peer merges all received responses and presents them to Dicoogle user.
If the peer X wants to retrieve an object DICOM, it uses a File Request message with the object FileName and FileHash, previously returned by Query Responses (Fig. 6 ). If the object is available on distinct peers, the request can be sent to multiple peers. Those peers will split the DICOM object in chunks, every one returned to peer X through a File Response message. The File Request message can request distinct DICOM objects, including still images, cine-loops, waveforms, structured reports, etc.
). If the object is available on distinct peers, the request can be sent to multiple peers. Those peers will split the DICOM object in chunks, every one returned to peer X through a File Response message. The File Request message can request distinct DICOM objects, including still images, cine-loops, waveforms, structured reports, etc.
In WAN communications, the relay receives the messages of the source peers and stores them. In the meantime, all network peers are constantly checking if there are new messages destined to them. After delivering the message to destination peer(s), it will be deleted from memory. To maximize the potential of the platform chosen (Google App Engine), we developed a multi-threaded mechanism, capable of increase or decrease the number of polling requests, accordingly to the number of messages in the relay. In other words, if the relay has many messages for the peer, the peer sends multiple requests to the cloud, in order to receive them in as fast as possible.
Security
This Dicoogle network protocol defines security domains through the concept of community or affinity group. One same relay service can support multiple groups and its administrator has access to a web interface, where it is possible to associate peers to domains. In this way, the traffic generated by a peer is only forwarded if the destination belongs to the same community. Therefore, the system allows the creation of federations of institutions capable of sharing their DICOM persistent objects.
To have access to a relay service, it is necessary to provide a username and password. After the login is carried out, the Dicoogle instance is able to receive and send messages to the community. A client keep-alive message is sent, at intervals of 30 seconds, to inform relay service that the peer is still active.
One issue in sending medical imaging data through Internet is data privacy. For that reason, to assure data privacy, the system uses HTTPS between peers and GAE. Moreover, affinity groups are identified by a farmID, a crypto token generated by the relay administrator. It is a 256 bits key used to assure communication’s privacy between peers, through Advanced Encryption Standard (AES) ciphering.
All Dicoogle network messages are ciphered with a farmID key. Those ciphered messages are encapsulated in HTTPS messages. The HTTP message header is also used to transmit some important data to forward medical imaging message, such as the name of the destination, the source peers and the message purpose.
RESULTS
Application
The proposed framework was integrated in Dicoogle Open Source project and is available for medical and scientific community at www.dicoogle.com, including binaries and source code. Consequently, the article results are focused on Dicoogle application that, implicitly, uses the proposed framework to support network processes. Fig. (7 ) presents the GUI of Dicoogle Desktop version. With this interface, users can select the range of searches performed (Fig. 7
) presents the GUI of Dicoogle Desktop version. With this interface, users can select the range of searches performed (Fig. 7 -1
-1 ). For instance, it can search on all interfaces at the same time. The results are presented following the DIM structure and organization (Patient/Studies/Series/Images), identifying the objects data sources (Fig. 7
). For instance, it can search on all interfaces at the same time. The results are presented following the DIM structure and organization (Patient/Studies/Series/Images), identifying the objects data sources (Fig. 7 -2
-2 ). If the object is in an external data source (LAN or WAN), the user can download it (Fig. 7
). If the object is in an external data source (LAN or WAN), the user can download it (Fig. 7 -3
-3 ). It is also possible to see the number of patients and objects returned (Fig. 7
). It is also possible to see the number of patients and objects returned (Fig. 7 -4
-4 ) from query. Finally, it is possible to see the list of active peers (and users) in affinity group (Fig. 7
) from query. Finally, it is possible to see the list of active peers (and users) in affinity group (Fig. 7 -5
-5 ).
).
This medical imaging platform has been used in several distributed environments. For instance, Dicoogle is used to support a regional PACS archive, integrating two diagnostic centres, with Radiologists reporting from several places [38Silva LAB, Costa C, Oliveira JL, Eds. An agile framework to support distributed medical imaging scenarios. IEEE International Conference on Healthcare Informatics 2013 (ICHI 2013) Philadelphia USA2013. ]. The load over the repository and network is considerable because the system manages more than 106k DICOM object per month. On other use case, Dicoogle is being used has DICOM data mining over several distributed repositories of hospitals from Aveiro region [8Santos M, Bastião L, Costa C, Silva A, Rocha N. DICOM and Clinical Data Mining in a Small Hospital PACS A Pilot Study.ENTER prise Information Systems. In: Cruz-Cunha M, Varajão J, Powell P, Martinho R, Eds. Communications in Computer and Information Science 221. Berlin Heidelberg;: Springer 2011; pp. 254-63.]. There is a Dicoogle installation in every hospital and it is possible to perform distributed queries over those federated indexes, representing more than 8 million of DICOM objects. The tool made possible to collect, identify and characterize DICOM metadata that can be related with different quality assurance vectors. The characterization of data inconsistencies and the identification of missing data, along with the collection of important data for practice improvement, are contributing for a better dose surveillance and patient safety.
Other innovator scenarios were potentiated with Dicoogle networking. As an example, Dicoogle Mobile [39Viana-Ferreira C, Ferreira D, Valente F, Monteiro E, Costa C, Oliveira J. Dicoogle Mobile a medical imaging platform for Android. Health Technol Inform 2012; 180: 502.] is an application developed for Android mobile devices. It uses the relay service to communicate with normal Dicoogle instances. In this way, this application is able to search and display medical images stored in other Dicoogle instances.
Performance Trials
To evaluate the performance of data search and results integration, especially in inter-institutional processes supported by Dicoogle private protocol, sometests were performed. For what concerns WAN testing, the bandwidth of Internet access from inside an institution depends on numerous factors like, for instance, the traffic generated by the users in a specific instant. Thus, tests were done to assess the average Internet access bandwidth of trials region, the results pointed to 17 megabits per second. For that reason, in the system’s tests, the peers had an Internet access bandwidth of 17 megabits per second of upstream and downstream. The LAN scenario was tested with 4 machines linked through a connection with a bandwidth of 100 megabits per second and with several machines generating some daily traffic. The peer, in which the local evaluation was carried out, was endowed with an IntelCoreDuo E8400 processor and 4GB of memory. Theother3machinesusedduring the tests were similar, or slightly worse than this one. In each machine we installed a Dicoogle peer application with medical imaging studies.
For each of those scenarios, numerous queries were sent, in order to assess the behaviour of the network under different distributions and numbers of peers. Fig. (8 ) shows the time length from the application sending the search request until the retrieval of results, in four main scenarios: (1) Only the local database is consulted (Local); (2) The peers consulted are in the same LAN, via DICOM (LAN 2 Peers, LAN 3 Peers, LAN 4 Peers); (3) The peers consulted are in the WAN, connected via Dicoogle GAE relay (WAN 2 Peers, WAN 3 Peers, WAN 4 Peers); (4) The peer that starts the search sends the query via WAN, this query is propagated also via DICOM to other peers not directly connected via WAN as depicted in Fig. (9
) shows the time length from the application sending the search request until the retrieval of results, in four main scenarios: (1) Only the local database is consulted (Local); (2) The peers consulted are in the same LAN, via DICOM (LAN 2 Peers, LAN 3 Peers, LAN 4 Peers); (3) The peers consulted are in the WAN, connected via Dicoogle GAE relay (WAN 2 Peers, WAN 3 Peers, WAN 4 Peers); (4) The peer that starts the search sends the query via WAN, this query is propagated also via DICOM to other peers not directly connected via WAN as depicted in Fig. (9 ), (a) is named as WAN2+LAN3, while (b) is named as WAN3+LAN2. In Fig. (9
), (a) is named as WAN2+LAN3, while (b) is named as WAN3+LAN2. In Fig. (9 ), the peer that starts the search is the one signalled as client and all the other peers collaborate in the search.
), the peer that starts the search is the one signalled as client and all the other peers collaborate in the search.
As expected, the local search is the fastest kind of search, being able to retrieve search results at a velocity that remains almost constant, independently of the number of search hits, at about 11000 results per second. On the other hand, the slowest ones are the hybrid ones. This was expected, due to the location and communication latency involved in this kind of search.
CONCLUSION
Medical imaging repositories are often looked on as “inert bags” of DICOM objects, accessible only though the DICOM query and retrieve service. In this paper, we have presented a framework capable of retrieving data from distinct sources through three communication protocols: XDS-I, DICOM and a private protocol based on P2P paradigm. As a proof-of-concept, we used Dicoogle that is an open source PACS solution with an extensible platform.
As a result, we have obtained a solution that can retrieve data from medical imaging repositories through distinct ways. Due to the variety of protocols used, the solution combines the advantages of the distinct paradigms, being easy to deploy and maintain, transparent and scalable.
The data source bus endows the system with the ability to give access to repositories that are not directly accessible, using a member of the network with direct access to the repository as an intermediary. In its turn, the private protocol gives to the system the ability to take the most of the complex searching mechanisms of the Dicoogle project. Nevertheless, the framework also supports standard protocols like DICOM and XDS-I to promote the interoperability to other components of healthcare institutions’ networks.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
This work has received funding from “Fundaçãopara a Ciência e Tecnologia” (FCT) under grant agreement PTDC/EIA-EIA/104428/2008. Carlos Viana-Ferreira is funded by the FCT grant SFRH/BD/68280/2010.
1 https://www.oasis-open.org/
2 ftp://ftp.ihe.net/TF_Implementation_Material/ITI/wsdl



