- Home
- About Journals
-
Information for Authors/ReviewersEditorial Policies
Publication Fee
Publication Cycle - Process Flowchart
Online Manuscript Submission and Tracking System
Publishing Ethics and Rectitude
Authorship
Author Benefits
Reviewer Guidelines
Guest Editor Guidelines
Peer Review Workflow
Quick Track Option
Copyediting Services
Bentham Open Membership
Bentham Open Advisory Board
Archiving Policies
Fabricating and Stating False Information
Post Publication Discussions and Corrections
Editorial Management
Advertise With Us
Funding Agencies
Rate List
Kudos
General FAQs
Special Fee Waivers and Discounts
- Contact
- Help
- About Us
- Search

The Open Anatomy Journal
(Discontinued)
ISSN: 1877-6094 ― Volume 6, 2014
Evaluation of the Microanatomy of the Liver via a Rapid Sample Preparation Protocol and a Table-Top Scanning Electron Microscope
Filip Braet*
Abstract
In this paper an alternative preparation and imaging method is presented that allows investigation of the fine structure of hepatic tissue within one hour after the initial fixation step. This approach involves, besides traditional fixation with glutaraldehyde, chemical drying and subsequent investigation of the samples with the new generation of user-friendly desktop scanning electron microscopes. The data presented herein reveal comparative preservation of liver ultrastructure to those that are prepared using classical sample preparation protocols. This low-cost approach permits the swift study of liver tissue subjected to various testing conditions and hence bridges the time gap between experiment and subsequent ultrastructural observation at the nanoscale.
Article Information
Identifiers and Pagination:
Year: 2010Volume: 2
First Page: 98
Last Page: 101
Publisher Id: TOANATJ-2-98
DOI: 10.2174/1877609401002010098
Article History:
Received Date: 05/11/2010Revision Received Date: 23/11/2010
Acceptance Date: 26/11/2010
Electronic publication date: 30/12/2010
Collection year: 2010
open-access license: This is an open access article licensed under the terms of the Creative Commons Attribution Non-Commercial License (http: //creativecommons.org/licenses/by-nc/3.0/ which permits unrestricted, non-commercial use, distribution and reproduction in any medium, provided the work is properly cited.
* Address correspondence to this author at the Australian Centre for Microscopy and Microanalysis (ACMM), Madsen Building F09, The University of Sydney, NSW 2006, Australia; Tel: + 61 2 9351 7619; Fax: + 61 2 9351 7682; E-mailfilip.braet@sydney.edu.au
| Open Peer Review Details | |||
|---|---|---|---|
| Manuscript submitted on 05-11-2010 |
Original Manuscript | Evaluation of the Microanatomy of the Liver via a Rapid Sample Preparation Protocol and a Table-Top Scanning Electron Microscope | |
INTRODUCTION
Nowadays the quest is on for alternative and quick sample preparation protocols to serve the demands of the impatient researcher and or to present a weight against the booming availability of fast cryo-sample preparation methods for electron microscopic investigation [1]. However, fast tracking classical sample preparation methods often result in the presence of structural artefacts which make an accurate assessment on the ultrastructure of the sample difficult [2]. This often leads to misleading morphological information, and hence conclusions [3, 4].
In this contribution an alternative quick approach is presented that allows investigation of the fine structure of hepatic tissue within one hour after the initial glutaraldehyde-fixation step. Subsequent steps involve the immediate dehydration and air drying of the sample by evaporation of hexamethyldisilazane [5, 6], which was immediately followed by sample mounting and sputter coating of the tissue blocks with a thin layer of gold [7]. This sample preparation method was combined with the new-generation of table-top scanning electron microscopes. These instruments are typically characterised by their ease of use and are within the budget of every research laboratory that endeavors to explore the microanatomy of tissues and cells over micro- to nanometer scales.
The data presented herein reveal comparative preservation of ultrastructure to those that are prepared using the well-described classical combined glutaraldehyde-osmium fixation method [8, 9], which is typically followed by time-consuming dehydration and subsequent critical point drying procedures. Furthermore, the method presented in this paper result in quality electron microscopic data at low and intermediate magnifications. This magnification range is often sufficient to address the majority of the research quests.
MATERIALS & METHODS
Tissue Preparation. The liver of eight weeks old male Sprague Dawley rats (n=4) was perfuse-fixed at physiological pressure (12 cm H20) with 1.5% glutaraldehyde in 0.1M Na-cacodylate buffer with 0.1 M sucrose for 2 min at room temperature. See the paper by Wisse & Braet et al. [10] for full experimental details regarding the perfusion-fixation procedure. Next, the liver was cut into small pieces of 1×1×1 mm with the aid of razor blade under 0.1M Na-cacodylate buffer supplemented with 0.1 M sucrose, in order to allow full immersion with the dehydration vehicle (see next step). The samples were next rinsed five times for 2 min each in hexamethyldisilazane followed by hexamethyldisilazane immersion for 10 min [7]. Subsequently, the tissue blocks were transferred to a dessicator and allowed to air-dry for 10 min. After drying, samples were immediately broken in half with a pair of tweezers, mounted on SEM stubs using conductive carbon tape and sputter-coated with 10 nm gold [7]. The different preparation steps are schematically depicted under Fig. (1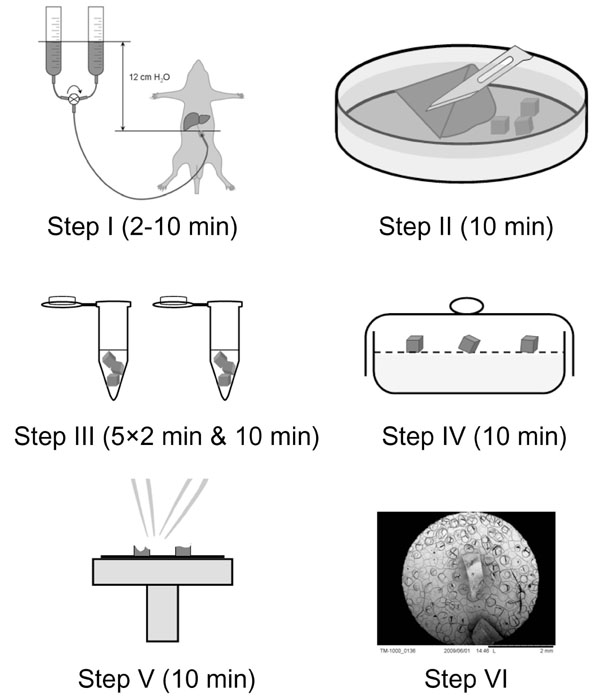 ).
).
Scanning Electron Microscopy. Samples were investigated in eucentric position at 15 kV using the TM-1000 Hitachi Tabletop Microscope (Hitachi Technologies, Japan). Digital images were captured using the auto image adjustment function running under Hitachi TM-1000 software (v1.01) interface. High-resolution data without digital zoom function were recorded at 1,280 × 1,040 dpi and at a working distance between 5.3 and 6.7 micrometers.
RESULTS & DISCUSSION
Following the steps as outlined under Fig. (1 ), allowed the observation of the microanatomy of liver tissue within an hour after the start of the perfusion-fixation process. Examination of the tissue blocks at low magnification (Fig. 2A
), allowed the observation of the microanatomy of liver tissue within an hour after the start of the perfusion-fixation process. Examination of the tissue blocks at low magnification (Fig. 2A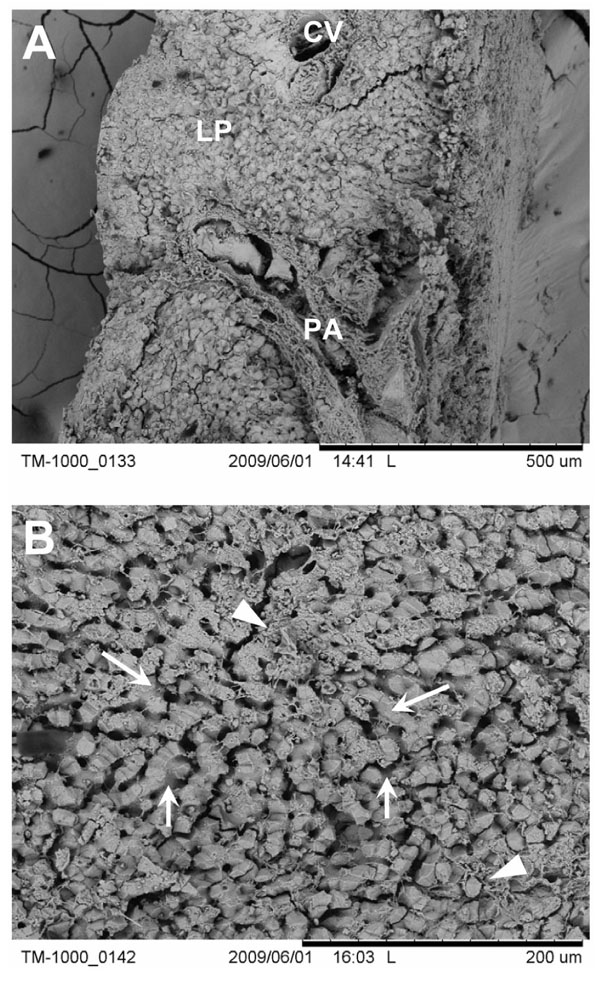 ) resulted in the overall observation of the histological architecture of the functional liver unit [11]. Large portal areas containing branches of the vena porta, hepatic artery and bile duct could be easily descerned together with the central veins that collect the blood that flow through the microvascular bed of the liver (i.e., the liver sinusoids). At intermediate magnification, the individual sinusoids were readily visible and were surrounded by the liver parenchymal tissue that was composed of hepatocytes (Fig. 2B
) resulted in the overall observation of the histological architecture of the functional liver unit [11]. Large portal areas containing branches of the vena porta, hepatic artery and bile duct could be easily descerned together with the central veins that collect the blood that flow through the microvascular bed of the liver (i.e., the liver sinusoids). At intermediate magnification, the individual sinusoids were readily visible and were surrounded by the liver parenchymal tissue that was composed of hepatocytes (Fig. 2B ). Also, fine extracellular matrix material such as fine collagen bundles could be easily noted.
). Also, fine extracellular matrix material such as fine collagen bundles could be easily noted.
High-magnification imaging of the areas around the microvascular bed permitted the observation of the individual hepatocytes that contained numerous microvillous protrusions. Furthermore, bile canaliculi could be easily seen running between adjacent hepatocytes (Fig. 3A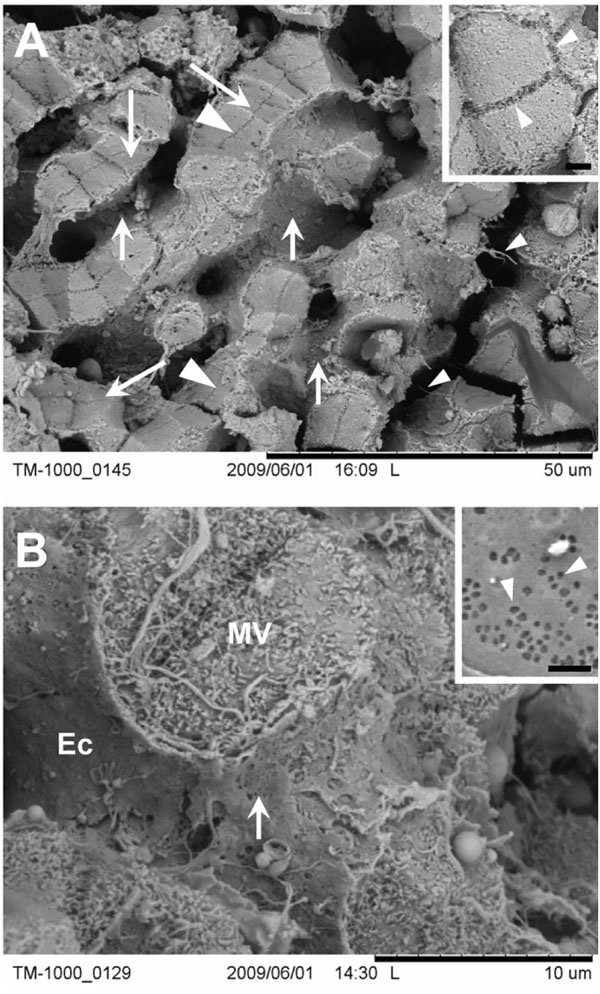 ). The table-top scanning electron microscopes allow magnification of samples up to 7,500-10,000 times. At this magnification range, the sinusoids disclosed their fenestrations, which have a typical size in the order of 120-160 nm (Fig. 3B
). The table-top scanning electron microscopes allow magnification of samples up to 7,500-10,000 times. At this magnification range, the sinusoids disclosed their fenestrations, which have a typical size in the order of 120-160 nm (Fig. 3B ). The dimension of this unique ultrastructural feature directly demonstrates the magnification capabilities of these types of table-top scanning electron microscopes.
). The dimension of this unique ultrastructural feature directly demonstrates the magnification capabilities of these types of table-top scanning electron microscopes.
The presence of fenestrae is also a direct structural indicator that the preparation method herein outlined is apparently sufficient enough for their preservation. This comes not as a surprise, since brief glutaraldehyde-fixation combined with cryo-fixation allowed the direct visualization of this fragile structures [12]. Noteworthy, fenestrae are known to be difficult to preserve and often collapse and/or fuse as a result of improper fixation or drying procedures [13, 14]. In line, drying of fenestrated liver endothelial cells via hexamethyldisilazane was found to be successful in the preservation of these fragile membrane-bound pores under in vitro conditions [7]. And drying by evaporation of hexamethyldisilazane has been described as a good alternative for a variety of biological samples [5, 6]. It is well known that air-drying of biological samples in water is disastrous, because of the destructive action of the surface tension during drying. Hexamethyldisilazane, however, has a reduced surface tension and also cross-links proteins, therefore adding strength to the sample during air-drying. This is supposed to reduce fracturing and collapsing of the specimen [15, 16]. All of this is in direct support of the present observation on liver tissue whereby hexamethyldisilazane does not affect the integrity of fine structural details.
In conclusion, a fast sample preparation method for the preservation of liver tissue for subsequent scanning electron microscopy investigation is herein presented. This method allows the facile collection of intermediate- up to high magnification electron microscopy data within an hour after the initial preparation step of liver tissue. Fine ultrastructural details within the nanometer regime can be easily retrieved from samples which are directly comparable to data obtained from samples prepared with conventional methods and imaging techniques, which requires more time and effort, including the cost of ‘high-end’ microscopes. This approach is therefore well-suited for implementation in basic and clinical settings in which time between experimentation and observation is central in the guidance to successive experimental testing thereby limiting the number of laboratory animals. In addition, the investigation of liver tissue after pharmacodynamic drug screening studies in animals or in the diagnostic examination of liver biopsies obtained from patients in the operating theatre would largely benefit from this approach.
Finally, it is noteworthy to flag that excellent standard sample preparation and scanning electron imaging methods do exist for the fine structure observation of liver tissue [9, 17]. These protocols have been proven successful at magnifications up to 30,000× and higher, and are without any doubt the first method of choice when the highest reachable resolution wants to achieved. This is particularly the case in field-emission scanning electron microscopy investigations that permit imaging down to one nanometer resolution.
ACKNOWLEDGEMENTS
The author is indebted is to Hitachi - Meeco Holdings Pty Ltd Australia and to Dr Joanna Biazik for reading of this manuscript.



