- Home
- About Journals
-
Information for Authors/ReviewersEditorial Policies
Publication Fee
Publication Cycle - Process Flowchart
Online Manuscript Submission and Tracking System
Publishing Ethics and Rectitude
Authorship
Author Benefits
Reviewer Guidelines
Guest Editor Guidelines
Peer Review Workflow
Quick Track Option
Copyediting Services
Bentham Open Membership
Bentham Open Advisory Board
Archiving Policies
Fabricating and Stating False Information
Post Publication Discussions and Corrections
Editorial Management
Advertise With Us
Funding Agencies
Rate List
Kudos
General FAQs
Special Fee Waivers and Discounts
- Contact
- Help
- About Us
- Search

The Open Virology Journal
(Discontinued)
ISSN: 1874-3579 ― Volume 15, 2021
Characterization of Alternanthera mosaic virus and its Coat Protein
Anna A Mukhamedzhanova*, 1, Alexander A Smirnov 1, Marina V Arkhipenko 1, Peter A Ivanov 1, Sergey N Chirkov 1, Nina P Rodionova 1, †, Olga V Karpova 1, Joseph G Atabekov 1, 2
Abstract
A new isolate of Alternantheramosaic virus (AltMV-MU) was purified from Portulaca grandiflora plants. It has been shown that the AltMV-MU coat protein (CP) can be efficiently reassembled in vitro under different conditions into helical RNA-free virus-like particles (VLPs) antigenically related to native virus. The AltMV-MU and VLPs were examined by atomic force and transmission electron microscopies. The encapsidated AltMV-MU RNA is nontranslatable in vitro. However, it can be translationally activated by CP phosphorylation or by binding to the TGB1protein from the virus-coded movement triple gene block.
Article Information
Identifiers and Pagination:
Year: 2011Volume: 5
First Page: 136
Last Page: 140
Publisher Id: TOVJ-5-136
DOI: 10.2174/1874357901105010136
Article History:
Received Date: 5/7/2011Revision Received Date: 9/9/2011
Acceptance Date: 21/9/2011
Electronic publication date: 21/11/2011
Collection year: 2011
open-access license: This is an open access article licensed under the terms of the Creative Commons Attribution Non-Commercial License (http: //creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted, non-commercial use, distribution and reproduction in any medium, provided the work is properly cited.
* Address correspondence to this author at the Department of Virology, Moscow State University, Moscow 119991, Russia; Tel: +7 495 939 33 47; Fax: +7 495 938 0601; E-mail: amukhamedzhanovamsu@gmail.com† Deceased on 14 April 2011.
| Open Peer Review Details | |||
|---|---|---|---|
| Manuscript submitted on 5-7-2011 |
Original Manuscript | Characterization of Alternanthera mosaic virus and its Coat Protein | |
Flexible filamentous virions of Alternanthera mosaic virus (AltMV) are typical for the family Flexiviridae (genus Potexvirus) [1Martelli GP, Adams MJ, Kreuze JF, Dolja VV. Family Flexiviridae: A case study in virion and genome plasticity Annu Rev Phytopathol 2007; 45: 73-100.]. AltMV was purified from Alternanthera pungenes (Amaranthaceae) in Australia by Geering and Thomas [2Geering AD, Thomas JE. Characterisation of a virus from Australia that is closely related to papaya mosaic potexvirus Arch Virol 1999; 144: 577-92.] who showed that AltMV was closely related to Papaya mosaic virus (PapMV). A potexvirus from symptomless portulaca plants (AltMV-Po) has been also isolated and characterized in Italy [3Ciuffo M, Turina M. A potexvirus related to Papaya mosaic virus isolated from moss rose (Portulaca grandiflora) in Italy Plant Pathol 2004; 53: 515.]. Furthermore, the AltMV isolates from phlox (AltMV-PA) and portulaca were identified in the United States [4Hammond J, Reinsel MD, Maroon-Lango CJ. Identification and full sequence of an isolate of Alternanthera mosaic potexvirus infecting Phlox stolonifera Arch Virol 2006; 151: 477-93.]. Recently, the complete nucleotide sequence of the genome of another AltMV isolate (AltMV-MU) has been determined by our group and deposited in GenBank (accession number FJ822136) [5Ivanov PA, Mukhamedzhanova AA, Smirnov AA, Rodionova NP, Karpova OV, Atabekov JG. The complete nucleotide sequence of Alternanthera mosaic virus infecting Portulaca grandi?ora represents a new strain distinct from phlox isolates Virus Genes 2011; 42: 268-71.]. The RNA sequence analysis allowed to characterize AltMV-MU as a separate strain with a new genotype. In the present work some properties of the virions and the coat protein (CP) of AltMV-MU that caused symptomless infection in Portulaca grandiflora plants were investigated.
TRANSMISSION ELECTRONIC AND ATOMIC FORCE MICROSCOPY OF VIRIONS AND VIRUS-LIKE PARTICLES FORMED BY ALTMV-MU COAT PROTEIN
The AltMV-MU has been propagated in P. grandiflora and purified from systemically infected leaves by the method developed for potato virus X (PVX) purification [6Atabekov JG, Rodionova NP, Karpova OV, Kozlovsky SV, Novikov VK, Arkhipenko MV. Translational activation of encapsidated potato virus X RNA by coat protein phosphorylation Virology 2001; 286: 466-74.] using a modified (50mM Tris-HCl, 10mM EDTA pH 8.0) extraction buffer. Virus concentration was estimated using an average extinction coefficient of 2.84 for potexviruses [2Geering AD, Thomas JE. Characterisation of a virus from Australia that is closely related to papaya mosaic potexvirus Arch Virol 1999; 144: 577-92.]. Purified virus had an A260/A280 ratio of 1.44 and yield was of 0.34 mg/g of leaves harvested at 21-25 days after inoculation.
Viral RNA was isolated from the purified viral particles by phenol deproteinization. The method of salt deproteinization with 2M LiCl described for PVX CP [7Homer RB, Goodman RM. Circular dichroism and fluorescence studies on potato virus X and its structural components Biochim Biophys Acta 1975; 378: 296-304.] was used to obtain the AltMV CP preparation. The extinction coefficient of 0.7 for AltMV CP was calculated (http://us.expasy.org/tools/protparam.html).
A single protein band with apparent size of 22 kDa was observed when dissociated virions and purified CP preparations were analyzed by 8-20% SDS-PAGE and Western blot (Fig. 1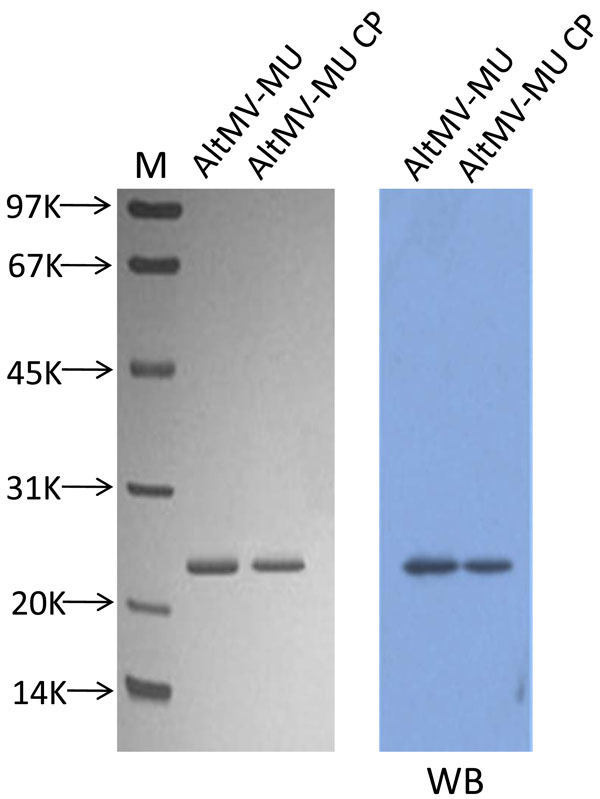 ). The Mr of the protein deduced from the amino acid sequence was 22, 1 x 103 which was close to the value determined by SDS-PAGE.
). The Mr of the protein deduced from the amino acid sequence was 22, 1 x 103 which was close to the value determined by SDS-PAGE.
For transmission electron microscopy (TEM) on copper grids [8Karpova OV, Zayakina OV, Arkhipenko MV, et al. Potato virus X RNA-mediated assembly of single tailed ternary “coat protein-RNA-movement protein” complexes J Gen Virol 2006; 87: 2731-40.], the purified viral particles (60 µg/ml) and CP aggregates (60 µg/ml) were negatively stained with 2% (w/v) uranyl acetate. Particles lengths were measured manually from digital prints by Image J program. Atomic force microscopy (AFM) measurement was carried out using a NanoScope IIIa multimode-scanning probe microscope (Digital Instruments) in tapping mode in air, as described previously [8Karpova OV, Zayakina OV, Arkhipenko MV, et al. Potato virus X RNA-mediated assembly of single tailed ternary “coat protein-RNA-movement protein” complexes J Gen Virol 2006; 87: 2731-40., 9Kiselyova OI, Yaminsky IV, Karpova OV, et al. AFM study of potato virus X disassembly induced by movement protein J Mol Biol 2003; 332: 321-5.].
According to the TEM data, the mean length and diameter of AltMV-MU virion were 570 nm and 13 nm, respectively (Fig. 2a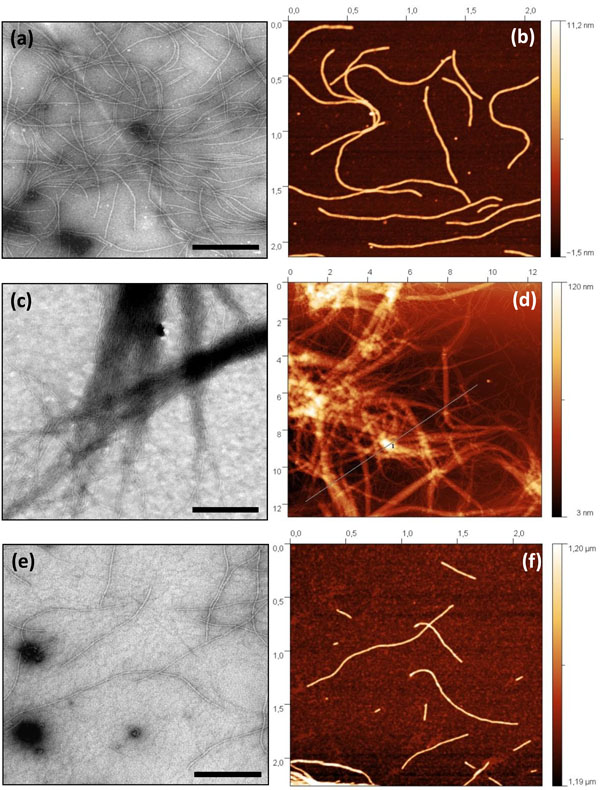 ). Based on AFM images, the virions average height was 6-8 nm (Fig. 2b
). Based on AFM images, the virions average height was 6-8 nm (Fig. 2b ).
).
The ability of PapMV CP to assemble VLPs in vitro at pH 4.0 in the absence of genomic RNA is known since seventies [10Erickson JW, Bancroft JB, Horne RW. The assembly of papaya mosaic virus protein Virology 1976; 72: 514-7.]. By contrast, assembly of PapMV CP at pH 8.0 was limited by production of “small 13-33S aggregates” [10Erickson JW, Bancroft JB, Horne RW. The assembly of papaya mosaic virus protein Virology 1976; 72: 514-7.]. Here, we examined the ability of RNA-free CP AltMV-MU to form VLPs under those conditions.
We found that similarly to PapMV CP, AltMV-MU CP could be assembled in vitro into VLPs at pH 4.0 and low ionic strength. In 0.01M citrate buffer, pH 4.0 a considerable part of AltMV-MU VLPs stuck together producing long bundles (Fig. 2c , d
, d ). However, in contrast to PapMV, TEM and AFM analyses showed that AltMV-MU CP formed filamentous VLPs at pH 8.0 as well (Fig. 2e
). However, in contrast to PapMV, TEM and AFM analyses showed that AltMV-MU CP formed filamentous VLPs at pH 8.0 as well (Fig. 2e , f
, f ). In 0.01M Tris-HCl buffer at pH 8.0, AltMV-MU CP formed VLPs, comparable in size to virus particles under the same condition (0.01M Tris-HCl buffer, pH 8.0) or even exceeding them in length (Fig. 2a
). In 0.01M Tris-HCl buffer at pH 8.0, AltMV-MU CP formed VLPs, comparable in size to virus particles under the same condition (0.01M Tris-HCl buffer, pH 8.0) or even exceeding them in length (Fig. 2a , b
, b , e
, e , f
, f ). In agreement with the AFM data the length of VLPs formed by AltMV-MU CP at pH 8.0 ranged from 60 nm to 2000 nm, the height was of 6-7 nm and the width of about 20 nm (Fig. 2f
). In agreement with the AFM data the length of VLPs formed by AltMV-MU CP at pH 8.0 ranged from 60 nm to 2000 nm, the height was of 6-7 nm and the width of about 20 nm (Fig. 2f ).
).
Apparently, the assembly of AltMV-MU CP into VLPs was less dependent on pH than that of PapMV. Therefore, the conditions of assembly and the size of the products formed from AltMV-MU and PapMV CPs were considerably different [10Erickson JW, Bancroft JB, Horne RW. The assembly of papaya mosaic virus protein Virology 1976; 72: 514-7.].
IMMUNOLOGICAL PROPERTIES OF VIRIONS AND VIRUS-LIKE PARTICLES
Antigenic specificity of AltMV-MU virions was compared with that of RNA-free VLPs assembled from AltMV-MU CP at pH 4.0. Antisera against AltMV-MU virions and VLPs have been obtained using a purified native virus preparation and a preparation of VLPs assembled at the pH 4.0 as antigens. Antigenic properties of AltMV-MU and VLPs were compared by indirect ELISA [11Clark MF, Adams AN. Characteristics of the microplate method of enzyme-linked immunosorbent assay for the detection of plant viruses J Gen Virol 1977; 34: 475-83.]. Purified preparations of native AltMV-MU and VLPs, both diluted with PBS pH 7.4 up to the concentration of 5 µg/ml, were applied to the wells of MaxiSorp microplate (Nunc) and adsorbed at 4°C overnight. Mouse antisera to AltMV-MU and VLPs as well as non-immune mouse serum used as a negative control were titrated against each antigen. Horseradish peroxidase-labelled anti-mouse IgG W402 (Promega) was used as detection antibodies.
Mouse antiserum to native AltMV-MU recognized the homologous antigen and some VLPs antigenic determinants which presumably are common for both structures but differently exposed on the VLPs and the native virus (Fig. 3a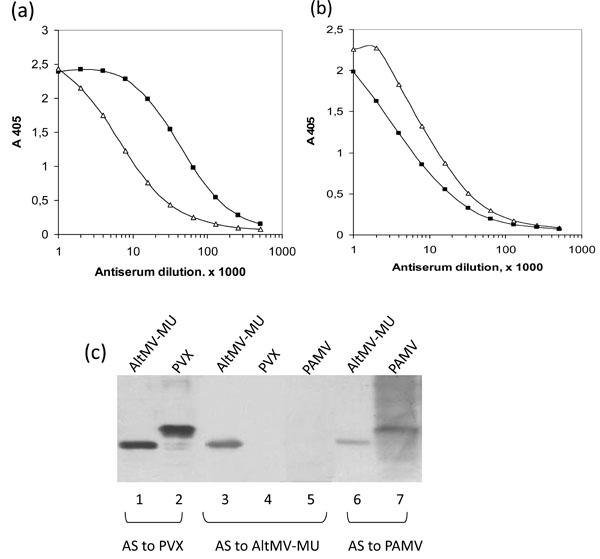 ). Similarly, mouse antiserum to VLPs strongly reacted with VLPs and also with native AltMV-MU even though the reaction with the virus was clearly weaker (Fig. 3b
). Similarly, mouse antiserum to VLPs strongly reacted with VLPs and also with native AltMV-MU even though the reaction with the virus was clearly weaker (Fig. 3b ). These data indicate that native AltMV-MU and VLPs are antigenically related, but not identical. This was not surprisingly because native virus particles (Fig. 2b
). These data indicate that native AltMV-MU and VLPs are antigenically related, but not identical. This was not surprisingly because native virus particles (Fig. 2b ) and VLPs assembled at the pH 4.0 in the absence of genomic RNA (Fig. 2d
) and VLPs assembled at the pH 4.0 in the absence of genomic RNA (Fig. 2d ) looked quite different morphologically suggesting some differences in their antigenic properties. In particular, some CP epitopes exposed at the surface of native virus particle, could be found hidden or disrupted at the VLPs assembly. On the other side, the neotope appearance cannot be ruled out at the VLPs formation.
) looked quite different morphologically suggesting some differences in their antigenic properties. In particular, some CP epitopes exposed at the surface of native virus particle, could be found hidden or disrupted at the VLPs assembly. On the other side, the neotope appearance cannot be ruled out at the VLPs formation.
The AltMV-MU, PVX and Potato aucuba mosaic virus (PAMV), members of Potexvirus group were analyzed by Western blotting. Blots were developed with polyclonal antisera against PVX diluted 1:20000, AltMV-MU - 1:10000 and PAMV - 1:3000 (Fig. 3c ). Both antisera to PVX and PAMV have been found to react with the AltMV-MU (Fig. 3c
). Both antisera to PVX and PAMV have been found to react with the AltMV-MU (Fig. 3c , lanes 1, 6). At the same time, the antiserum to AltMV-MU reacted neither with PVX nor with PAMV (Fig. 3c
, lanes 1, 6). At the same time, the antiserum to AltMV-MU reacted neither with PVX nor with PAMV (Fig. 3c , lanes 4, 5). Phylogenetic analysis of 26 coat protein sequences from genus Potexvirus was performed previously [5Ivanov PA, Mukhamedzhanova AA, Smirnov AA, Rodionova NP, Karpova OV, Atabekov JG. The complete nucleotide sequence of Alternanthera mosaic virus infecting Portulaca grandi?ora represents a new strain distinct from phlox isolates Virus Genes 2011; 42: 268-71.]. It shows that all three proteins are distanly located from each other within the appropriate tree that might explain the absence of cross-reactivity in case of antiserum to AltMV-MU.
, lanes 4, 5). Phylogenetic analysis of 26 coat protein sequences from genus Potexvirus was performed previously [5Ivanov PA, Mukhamedzhanova AA, Smirnov AA, Rodionova NP, Karpova OV, Atabekov JG. The complete nucleotide sequence of Alternanthera mosaic virus infecting Portulaca grandi?ora represents a new strain distinct from phlox isolates Virus Genes 2011; 42: 268-71.]. It shows that all three proteins are distanly located from each other within the appropriate tree that might explain the absence of cross-reactivity in case of antiserum to AltMV-MU.
TRANSLATIONAL ACTIVATION OF ALTMV-MU VIRIONS BY TGB1 PROTEIN
Recently we have shown that, contrary to Tobacco mosaic virus (TMV) and some other viruses, encapsidated PVX RNA was completely nontranslatable in vitro. The translational activation of PVX virions may be reached by two ways: either phosphorylation of PVX CP or TGB1 (triple gene block 1) protein binding to terminal subunits of the polar PVX helix, TGB1 is one of three PVX movement proteins [6Atabekov JG, Rodionova NP, Karpova OV, Kozlovsky SV, Novikov VK, Arkhipenko MV. Translational activation of encapsidated potato virus X RNA by coat protein phosphorylation Virology 2001; 286: 466-74., 12Atabekov JG, Rodionova NP, Karpova OV, Kozlovsky SV, Poljakov VY. The movement protein-triggered in situ conversion of potato virus X virion RNA from a nontranslatable into a translatable form Virology 2000; 271: 259-63.]. It is believed that binding of TGB1 to one end of PVX particles induces a structural transition (remodeling) of the PVX CP into a metastable form. As a result of this transition, the 5' end of viral RNA becomes accessible for ribosomes [13Rodionova NP, Karpova OV, Kozlovsky SV, Zayakina OV, Arkhipenko MV, Atabekov JG. Linear remodeling of helical virus by movement protein binding J Mol Biol 2003; 333: 565-72.]. Here, we report that similarly to virion PVX RNA, AltMV-MU encapsidated RNA was also untranslatable in vitro (Fig. 4b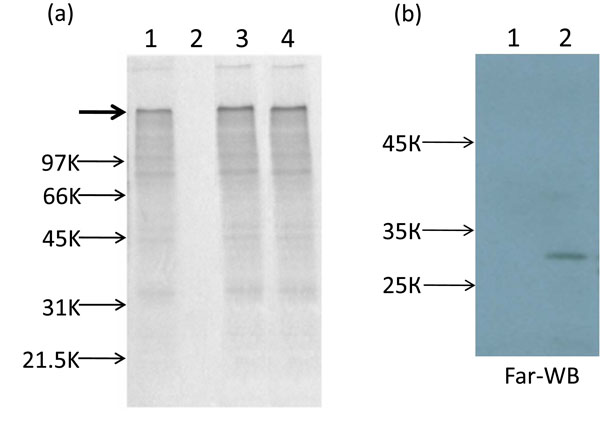 , lane 2) but could be activated by CP phosphorylation of the AltMV-MU virions by protein kinase C (PKC) (Fig. 4a
, lane 2) but could be activated by CP phosphorylation of the AltMV-MU virions by protein kinase C (PKC) (Fig. 4a , lane 3).
, lane 3).
A similar effect could be reached as a result of interaction of virions with AltMV-MU TGB1. The AltMV-MU TGB1 with N-terminal thioredoxin (Trx-tag), S-tag and His-tag was expressed in E. coli BL21 (DE3) strain using the pET 32(a)+ vector according to a Novagen protocol (Novagen pET System Manual, 11th Edition). The fusion protein with His-tag was purified using Ni-NTA affinity resin (Qiagen) and then specifically digested with enterokinase (Novagen, Tag-off rEK Cleavage/Capture Kit) to remove the Trx-tag, S-tag and His-tag. The data presented at Fig. (4a ) lane 4 showed that putative TGB1-CP AltMV-MU interaction resulted in translation of encapsidated genome RNA in vitro. This interaction was confirmed by Far-western blot analysis (Fig. 4b
) lane 4 showed that putative TGB1-CP AltMV-MU interaction resulted in translation of encapsidated genome RNA in vitro. This interaction was confirmed by Far-western blot analysis (Fig. 4b ). It could be presumed that two ways of translational activation of encapsidated RNA are peculiar features of some other members of genus Potexvirus.
). It could be presumed that two ways of translational activation of encapsidated RNA are peculiar features of some other members of genus Potexvirus.
It is known that the PapMV VLPs have been used successfully as an epitope presentation system. Moreover, PapMV VLP platforms have the advantage of triggering both arms of the immune responses, namely, the humoral and CTL responses, a characteristic that is unique for vaccine platforms [14Denis J, Majeau N, Acosta-Ramirez E, et al. Immunogenicity of papaya mosaic virus-like particles fused to a hepatitis C virus epitope: evidence for the critical function of multimerization Virology 2007; 363: 59-68., 15Leclerc D, Beauseigle D, Denis J, et al. Proteasome-independent major histocompatibility complex class I cross-presentation mediated by papaya mosaic virus-like particles leads to expansion of specific human T cells J Virol 2007; 81: 1319-26.]. As mentioned above, AltMV-MU is closely related to papaya mosaic virus (PapMV) [5Ivanov PA, Mukhamedzhanova AA, Smirnov AA, Rodionova NP, Karpova OV, Atabekov JG. The complete nucleotide sequence of Alternanthera mosaic virus infecting Portulaca grandi?ora represents a new strain distinct from phlox isolates Virus Genes 2011; 42: 268-71.]. We believe that AltMV-MU VLPs can be regarded as good candidates for presentation of foreign epitopes as well.
ACKNOWLEDGEMENTS
We are grateful to Dr. Stephan Winter (German Collection of Microorganisms and Cell Cultures, Braunschweig) for providing the initial portulaca isolate and Dr Sergey Saunin and Mikhail Savvateev (AIST-NT Company) for Atomic force microscopy assays. This work was funded in part by the Russian Foundation for Basic Research (Grant No. 10-04-000-89-a) and the Federal Agency of Science and Innovation (FASI) (contracts No 16.512.11.2184).






